Multivariate Analysis
Multidimensional Scaling
Multidimensional scaling is a collection of statistical techniques for exploring similarities and dissimilarities in
data. Multidimensional scaling takes item-item similarities and assigns each to a location in a low-dimensional space. In
this respect it is similar to other data reduction techniques, such as, factor analysis.
There are several variaites of multidimensional scaling; classical MDS, modern metric MDS, and modern nonmetric MDS.
As of Stata 10 there is support for classical MDS, modern metric MDS and nonmetric MDS. Whereas, Stata 9 only supported classical metric
multidimensional scaling for dissimilarity between observations. For the purposes of this introductory unit, we will
demonstrate multidimensional scaling using classical MDS.
Simple Example
This simple example uses airline distances between 10 US cities as the disimilarities.
/* enter distance matrix */
#delimit ;
matrix d = (
0,587,1212,701,1936,604,748,2139,2182,543\
587,0,920,940,1745,1188,713,1858,1737,597\
1212,920,0,879,831,1726,1631,949,1021,1494\
701,940,879,0,1374,968,1420,1645,1891,1220\
1936,1745,831,1374,0,2339,2451,347,959,2300\
604,1188,1726,968,2339,0,1092,2594,2734,923\
748,713,1631,1420,2451,1092,0,2571,2408,205\
2139,1858,949,1645,347,2594,2571,0,678,2442\
2182,1737,1021,1891,959,2734,2408,678,0,2329\
543,597,1494,1220,2300,923,205,2442,2329,0);
#delimit cr
global names atl chi den hou la mi ny sf sea dc
matrix rownames d = $names
matrix colnames d = $names
matrix list d
symmetric d[10,10]
atl chi den hou la mi ny sf sea dc
atl 0
chi 587 0
den 1212 920 0
hou 701 940 879 0
la 1936 1745 831 1374 0
mi 604 1188 1726 968 2339 0
ny 748 713 1631 1420 2451 1092 0
sf 2139 1858 949 1645 347 2594 2571 0
sea 2182 1737 1021 1891 959 2734 2408 678 0
dc 543 597 1494 1220 2300 923 205 2442 2329 0
mdsmat d, names($names)
Classical metric multidimensional scaling
dissimilarity matrix: d
Number of obs = 10
Eigenvalues > 0 = 6 Mardia fit measure 1 = 0.9954
Retained dimensions = 2 Mardia fit measure 2 = 1.0000
--------------------------------------------------------------------------
| abs(eigenvalue) (eigenvalue)^2
Dimension | Eigenvalue Percent Cumul. Percent Cumul.
-------------+------------------------------------------------------------
1 | 9582144.3 84.64 84.64 96.99 96.99
2 | 1686820.2 14.90 99.54 3.01 100.00
-------------+------------------------------------------------------------
3 | 8157.2984 0.07 99.61 0.00 100.00
4 | 1432.8699 0.01 99.63 0.00 100.00
5 | 508.66869 0.00 99.63 0.00 100.00
6 | 25.143486 0.00 99.63 0.00 100.00
--------------------------------------------------------------------------
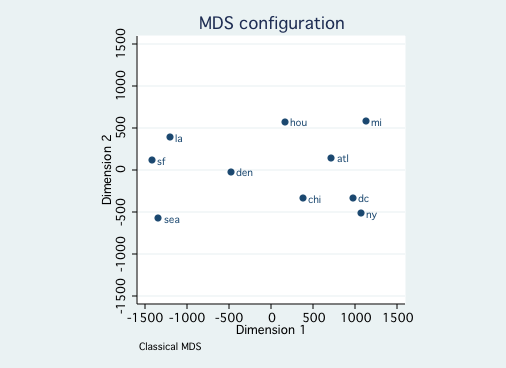 mdsconfig, autoaspect ynegate
mdsconfig, autoaspect ynegate
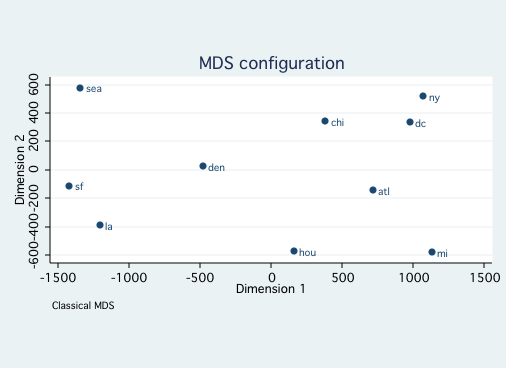 estat config
Approximating configuration in 2-dimensional Euclidean space
Category | dim1 dim2
-------------+----------------------------
atl | 718.7594 142.9943
chi | 382.0558 -340.8396
den | -481.6023 -25.2850
hou | 161.4663 572.7699
la | -1203.7380 390.1003
mi | 1133.5271 581.9073
ny | 1072.2357 -519.0242
sf | -1420.6033 112.5892
sea | -1341.7225 -579.7393
dc | 979.6220 -335.4728
------------------------------------------
estat config
Approximating configuration in 2-dimensional Euclidean space
Category | dim1 dim2
-------------+----------------------------
atl | 718.7594 142.9943
chi | 382.0558 -340.8396
den | -481.6023 -25.2850
hou | 161.4663 572.7699
la | -1203.7380 390.1003
mi | 1133.5271 581.9073
ny | 1072.2357 -519.0242
sf | -1420.6033 112.5892
sea | -1341.7225 -579.7393
dc | 979.6220 -335.4728
------------------------------------------
The next two examples are taken from the Stata manual.
Euclidean Distances Example
This dataset consists of eight variables with nutrition data on 25 breakfast cereals.
use http://www.stata-press.com/data/r9/cerealnut, clear
describe
Contains data from http://www.stata-press.com/data/r9/cerealnut.dta
obs: 25 Cereal Nutrition
vars: 9 24 Feb 2005 17:19
size: 1,150 (99.9% of memory free) (_dta has notes)
-------------------------------------------------------------------------------
storage display value
variable name type format label variable label
-------------------------------------------------------------------------------
brand str25 %25s Cereal Brand
calories int %9.0g Calories (Cal/oz)
protein byte %9.0g Protein (g)
fat byte %9.0g Fat (g)
Na int %9.0g Na (mg)
fiber float %9.0g Fiber (g)
carbs float %9.0g Carbs (g)
sugar byte %9.0g Sugar (g)
K int %9.0g K (mg)
-------------------------------------------------------------------------------
summarize calories-K
Variable | Obs Mean Std. Dev. Min Max
-------------+--------------------------------------------------------
calories | 25 109.6 21.30728 50 160
protein | 25 2.68 1.314027 1 6
fat | 25 .92 .7593857 0 2
Na | 25 195.8 71.32204 0 320
fiber | 25 1.7 2.056494 0 9
-------------+--------------------------------------------------------
carbs | 25 15.3 4.028544 7 22
sugar | 25 7.4 4.609772 0 14
K | 25 90.6 77.5043 15 320
replace brand = subinstr(brand, " ","_",.)
note the three variables, calories, Na, and K, which have standard deviations that are much
higher than the other variables.
The replace ckommand was used to replace spaces with underscores to make the graphs cleaner
and easier to interpret.
list brand, clean
brand
1. Cheerios
2. Cocoa_Puffs
3. Honey_Nut_Cheerios
4. Kix
5. Lucky_Charms
6. Oatmeal_Raisin_Crisp
7. Raisin_Nut_Bran
8. Total_Corn_Flakes
9. Total_Raisin_Bran
10. Trix
11. Wheaties_Honey_Gold
12. All-Bran
13. Apple_Jacks
14. Corn_Flakes
15. Corn_Pops
16. Mueslix_Crispy_Blend
17. Nut_&_Honey_Crunch
18. Nutri_Grain_Almond_Raisin
19. Nutri_Grain_Wheat
20. Product_19
21. Raisin_Bran
22. Rice_Krispies
23. Special_K
24. Life
25. Puffed_Rice
mds calories-K, id(brand) config
Classical metric multidimensional scaling
dissimilarity: L2, computed on 8 variables
Number of obs = 25
Eigenvalues > 0 = 8 Mardia fit measure 1 = 0.9603
Retained dimensions = 2 Mardia fit measure 2 = 0.9970
--------------------------------------------------------------------------
| abs(eigenvalue) (eigenvalue)^2
Dimension | Eigenvalue Percent Cumul. Percent Cumul.
-------------+------------------------------------------------------------
1 | 158437.92 56.95 56.95 67.78 67.78
2 | 108728.77 39.08 96.03 31.92 99.70
-------------+------------------------------------------------------------
3 | 10562.645 3.80 99.83 0.30 100.00
4 | 382.67849 0.14 99.97 0.00 100.00
5 | 69.761715 0.03 99.99 0.00 100.00
6 | 12.520822 0.00 100.00 0.00 100.00
7 | 5.7559984 0.00 100.00 0.00 100.00
8 | 2.2243244 0.00 100.00 0.00 100.00
--------------------------------------------------------------------------
Approximating configuration in 2-dimensional Euclidean space
brand | dim1 dim2
-------------+----------------------------
Cheerios | -61.8271 72.5534
Cocoa_Puffs | 38.5094 5.1037
Honey_Nut_~s | -28.0515 46.0667
Kix | 9.1693 81.4942
Lucky_Charms | 38.5024 5.1356
Oatmeal_Ra~p | -12.5635 -37.0897
Raisin_Nut~n | -12.0040 -73.7800
Total_Corn~s | 44.9827 33.2502
Total_Rais~n | -117.0067 -77.9962
Trix | 85.0033 -12.9330
Wheaties_H~d | 23.7367 19.7182
All-Bran | -226.1791 -67.6752
Apple_Jacks | 88.6199 -28.4323
Corn_Flakes | -1.8069 109.3770
Corn_Pops | 115.5366 -52.7072
Mueslix_Cr~d | -37.7449 -74.4727
Nut_&_Hone~h | 45.3886 21.9393
Nutri_Grai~n | -47.9441 0.6082
Nutri_Grai~t | 15.2261 -21.7290
Product_19 | -26.0875 129.4798
Raisin_Bran | -134.8587 -66.7255
Rice_Krisp~s | -2.3710 109.6115
Special_K | 12.1670 47.9540
Life | 20.9036 -41.4515
Puffed_Rice | 170.6994 -127.2995
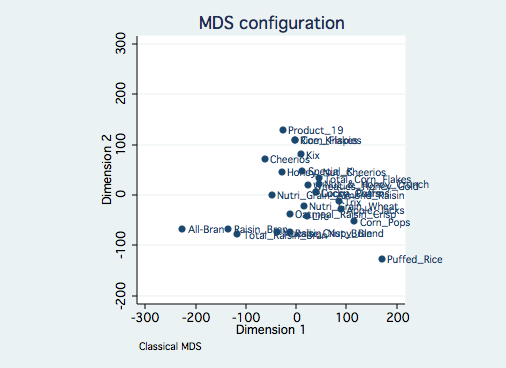
With a little bit of work we can make the graph clearer by making use of the mlabvopos
option to move the lable to different clock positions indicated by the new variable place.
generate place = 3
replace place = 9 if inlist(brand, "All_Bran","Corn_Flakes","Honey_Nut_Cheerios", ///
"Wheaties_Honey_Gold","Nutri_Grain_Wheat","Nutri_Grain_Almond_Raisin", ///
"Oatmeal_Raisin_Crisp","Cocoa_Puffs","Total_Raisin_Bran")
replace place = 6 if inlist(brand,"Mueslix_Crispy_Blend")
replace place = 12 if inlist(brand, "Raisin_Bran","Special_K")
mdsconfig, autoaspect mlabvpos(place)
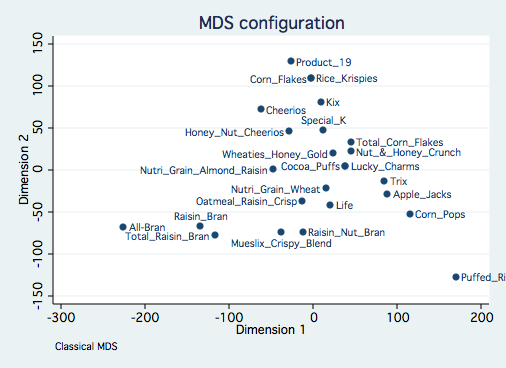
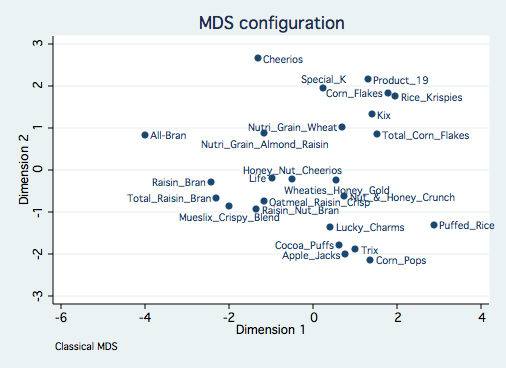
This configuration is due in large to the three variables with large standard deviations. An
alternative is to analyze the data with standardized variables using the std option. This
analysis with standardized Euclidean distances is equivalent to a principal components analysis
of the correlation among the variables.
mds calories-K, id(brand) config std noplot
Classical metric multidimensional scaling
dissimilarity: L2, computed on 8 variables
Number of obs = 25
Eigenvalues > 0 = 8 Mardia fit measure 1 = 0.5987
Retained dimensions = 2 Mardia fit measure 2 = 0.7697
--------------------------------------------------------------------------
| abs(eigenvalue) (eigenvalue)^2
Dimension | Eigenvalue Percent Cumul. Percent Cumul.
-------------+------------------------------------------------------------
1 | 65.645395 34.19 34.19 49.21 49.21
2 | 49.311416 25.68 59.87 27.77 76.97
-------------+------------------------------------------------------------
3 | 38.826608 20.22 80.10 17.21 94.19
4 | 17.727805 9.23 89.33 3.59 97.78
5 | 11.230087 5.85 95.18 1.44 99.22
6 | 8.2386231 4.29 99.47 0.78 99.99
7 | .77953426 0.41 99.87 0.01 100.00
8 | .24053137 0.13 100.00 0.00 100.00
--------------------------------------------------------------------------
Approximating configuration in 2-dimensional Euclidean space
brand | dim1 dim2
-------------+----------------------------
Cheerios | -1.3080 2.6638
Cocoa_Puffs | 0.6296 -1.7910
Honey_Nut_~s | -0.5050 -0.2227
Kix | 1.4003 1.3242
Lucky_Charms | 0.4178 -1.3534
Oatmeal_Ra~p | -1.1762 -0.7533
Raisin_Nut~n | -1.3523 -0.9414
Total_Corn~s | 1.5175 0.8541
Total_Rais~n | -2.3049 -0.6710
Trix | 1.0107 -1.8899
Wheaties_H~d | 0.5404 -0.2336
All-Bran | -4.0119 0.8411
Apple_Jacks | 0.7712 -2.0103
Corn_Flakes | 1.7864 1.8346
Corn_Pops | 1.3661 -2.1499
Mueslix_Cr~d | -2.0077 -0.8722
Nut_&_Hone~h | 0.7470 -0.6259
Nutri_Grai~n | -1.1706 0.8679
Nutri_Grai~t | 0.6929 1.0345
Product_19 | 1.3073 2.1645
Raisin_Bran | -2.4414 -0.2820
Rice_Krisp~s | 1.9619 1.7543
Special_K | 0.2362 1.9531
Life | -0.9843 -0.1881
Puffed_Rice | 2.8769 -1.3072
drop place
generate place = 3
replace place = 9 if inlist(brand, "All_Bran","Corn_Flakes", ///
"Nutri_Grain_Wheat","Apple_Jacks","Life", "Raisin_Bran" ///
"Oatmeal_Raisin_Crisp","Cocoa_Puffs","Total_Raisin_Bran")
replace place = 6 if inlist(brand,"Mueslix_Crispy_Blend","Nutri_Grain_Almond_Raisin", ///
"Wheaties_Honey_Gold")
replace place = 12 if inlist(brand,"Special_K","Honey_Nut_Cheerios")
mdsconfig, autoaspect mlabvpos(place)
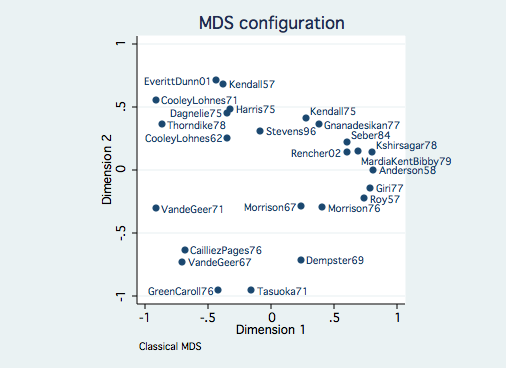
Non-Euclidean Dissimilarity Example
This dataset consists of seven variables measuring the number of topic pages
in 25 multivariate statistics books.
use http://www.stata-press.com/data/r9/mvstatsbooks, clear
describe
Contains data from http://www.stata-press.com/data/r9/mvstatsbooks.dta
obs: 25
vars: 8 15 Mar 2005 16:27
size: 825 (99.9% of memory free) (_dta has notes)
-------------------------------------------------------------------------------
storage display value
variable name type format label variable label
-------------------------------------------------------------------------------
author str17 %17s
math int %9.0g math other than statistics
(e.g., linear algebra)
corr int %9.0g correlation and regression,
including linear structural and
functional equations
fact byte %9.0g factor analysis and principal
component analysis
cano byte %9.0g canonical correlation analysis
disc int %9.0g discriminant analysis,
classification, and cluster
analysis
stat int %9.0g statistics, incl. dist. theory,
hypothesis testing & est.;
categorical data
mano int %9.0g manova and the general linear
model
-------------------------------------------------------------------------------
list, clean noobs
author math corr fact cano disc stat mano
Roy57 31 0 0 0 0 164 11
Kendall57 0 16 54 18 27 13 14
Kendall75 0 40 32 10 42 60 0
Anderson58 19 0 35 19 28 163 52
CooleyLohnes62 14 7 35 22 17 0 56
CooleyLohnes71 20 69 72 33 55 0 32
Morrison67 74 0 86 14 0 84 48
Morrison76 78 0 80 5 17 105 60
VandeGeer67 74 19 33 12 26 0 0
VandeGeer71 80 68 67 15 29 0 0
Dempster69 108 48 4 10 46 108 0
Tasuoka71 109 13 5 17 39 32 46
Harris75 16 35 69 24 0 26 41
Dagnelie75 26 86 60 6 48 48 28
GreenCaroll76 290 10 6 0 8 0 2
CailliezPages76 184 48 82 42 134 0 0
Giri77 29 0 0 0 41 211 32
Gnanadesikan77 0 19 56 0 39 75 0
Kshirsagar78 0 22 45 42 60 230 59
Thorndike78 30 128 90 28 48 0 0
MardiaKentBibby79 34 28 68 19 67 131 55
Seber84 16 0 59 13 116 129 101
Stevens96 23 87 67 21 30 43 249
EverittDunn01 0 54 65 0 56 20 30
Rencher02 38 0 71 19 105 135 131
mds math-mano, id(author) measure(corr) config noplot
Classical metric multidimensional scaling
similarity: correlation, computed on 7 variables
dissimilarity: sqrt(2(1-similarity))
Number of obs = 25
Eigenvalues > 0 = 6 Mardia fit measure 1 = 0.6680
Retained dimensions = 2 Mardia fit measure 2 = 0.8496
--------------------------------------------------------------------------
| abs(eigenvalue) (eigenvalue)^2
Dimension | Eigenvalue Percent Cumul. Percent Cumul.
-------------+------------------------------------------------------------
1 | 8.469821 38.92 38.92 56.15 56.15
2 | 6.0665813 27.88 66.80 28.81 84.96
-------------+------------------------------------------------------------
3 | 3.8157101 17.53 84.33 11.40 96.35
4 | 1.6926956 7.78 92.11 2.24 98.60
5 | 1.2576053 5.78 97.89 1.24 99.83
6 | .45929376 2.11 100.00 0.17 100.00
--------------------------------------------------------------------------
Approximating configuration in 2-dimensional Euclidean space
author | dim1 dim2
-------------+----------------------------
Roy57 | 0.7420 -0.2268
Kendall57 | -0.3794 0.6839
Kendall75 | 0.2763 0.4147
Anderson58 | 0.8144 -0.0001
CooleyLoh~62 | -0.3466 0.2573
CooleyLoh~71 | -0.9160 0.5594
Morrison67 | 0.2397 -0.2910
Morrison76 | 0.4088 -0.2976
VandeGeer67 | -0.7041 -0.7320
VandeGeer71 | -0.9173 -0.3062
Dempster69 | 0.2363 -0.7154
Tasuoka71 | -0.1549 -0.9563
Harris75 | -0.3220 0.4854
Dagnelie75 | -0.3509 0.4532
GreenCaro~76 | -0.4211 -0.9574
CailliezP~76 | -0.6827 -0.6365
Giri77 | 0.7898 -0.1441
Gnanadesi~77 | 0.3820 0.3612
Kshirsagar78 | 0.8014 0.1435
Thorndike78 | -0.8695 0.3641
MardiaKen~79 | 0.6923 0.1491
Seber84 | 0.6004 0.2254
Stevens96 | -0.0850 0.3124
EverittDu~01 | -0.4346 0.7139
Rencher02 | 0.6007 0.1399
------------------------------------------
generate spot = 3
replace spot = 2 if inlist(author,"Seber84","Kshirsagar78","Kendall75")
replace spot = 5 if author == "MardiaKentBibby79"
replace spot = 9 if inlist(author,"Dagnelie75","Rencher02", ///
"GreenCaroll76","EverittDunn01","CooleyLohnes62","Morrison67")
mdsconfig, mlabvpos(spot)
Multivariate Course Page
Phil Ender, 10may05
 mdsconfig, autoaspect ynegate
mdsconfig, autoaspect ynegate
 estat config
Approximating configuration in 2-dimensional Euclidean space
Category | dim1 dim2
-------------+----------------------------
atl | 718.7594 142.9943
chi | 382.0558 -340.8396
den | -481.6023 -25.2850
hou | 161.4663 572.7699
la | -1203.7380 390.1003
mi | 1133.5271 581.9073
ny | 1072.2357 -519.0242
sf | -1420.6033 112.5892
sea | -1341.7225 -579.7393
dc | 979.6220 -335.4728
------------------------------------------
estat config
Approximating configuration in 2-dimensional Euclidean space
Category | dim1 dim2
-------------+----------------------------
atl | 718.7594 142.9943
chi | 382.0558 -340.8396
den | -481.6023 -25.2850
hou | 161.4663 572.7699
la | -1203.7380 390.1003
mi | 1133.5271 581.9073
ny | 1072.2357 -519.0242
sf | -1420.6033 112.5892
sea | -1341.7225 -579.7393
dc | 979.6220 -335.4728
------------------------------------------


